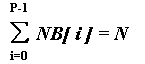
| C-DVM - contents | Part 1(1-4) | Part 2 (5-11) | Part 3 (Appendixes) |
| created: april, 2001 | - last edited 08.10.02 - |
C-DVM language was designed to support the development of portable and efficient parallel computational applications. The language is an extension of C language according to DVM model developed in Keldysh Institute of Applied Mathematics [1].
Traditionally applications for serial computers were created using Fortran and C languages. At present such applications are created for multiprocessor computers with distributed memory and networks using as a rule Fortran 77 and C extended by message transfer libraries (PVM, MPI). The development of such parallel applications requires from a programmer far more efforts than development of sequential ones, as the programmer has to distribute data and computations between different processors and also provide interaction of processors via message passing. Actually, the parallel application is a system of interacting programs, each program running at its own processor. The programs running at different processors may be either quite different, or may differ a little, or may be the same program whose behavior depends on the processor number. However, even in the last case the programmer is forced to develop and maintain two versions of the program - the sequential and parallel ones.
Using C-DVM language the programmer deals with the only version of the program both for sequential and parallel execution. Besides algorithm description by means of usual C features the program contains rules for parallel execution of the algorithm. These rules are syntactically organized in such a manner that they are "transparent" for standard C compilers at serial computers. Therefore it is possible to execute and to debug the C-DVM program at workstations as a usual sequential one.
Thus the C-DVM language is a standard C language extended by features to describe the rules of parallel program execution. The C-DVM language provides:
To create C-DVM programs the special system of parallel program development automation, named DVM-system, is provided.
DVM-system provides unified toolkit to develop parallel programs of scientific-technical calculations in C and Fortran 77.
DVM parallel model. DVM parallel model is based on data parallel model. The DVM name reflects two names of the model - Distributed Virtual Memory and Distributed Virtual Machine. These two names show that DVM model is adopted both for shared memory systems and for distributed memory systems. DVM high level model allows not only to decrease cost of parallel program development but provides unified formalized base for supporting Run-Time System, debugging, performance analyzing and prediction.
Languages and compilers. As distinct from HPF standard the goal of full automation of computation parallelization and the common data access synchronization is not assumed in DVM system. Using high-level specifications, a programmer has full control of parallel program performance.
Unified parallel model is built in C and Fortran 77 language on the base of the constructions that are "transparent" for standard compilers that allows to have single version of the program for sequential and parallel execution. C-DVM and Fortran DVM compilers translate DVM-program in C or Fortran 77 program correspondently, including parallel execution Run-Time Support system calls. So only requirement to a parallel system is availability of C and Fortran 77 compilers.
Execution and debugging technics. Unified parallel model allows to have unified Run-Time Support system for both languages and, as result, unified system of debugging, performance analyzing and prediction. There are following modes of DVM-program execution and debugging:
Following debugging modes are provided in pseudo-parallel and parallel execution modes:
2.1 Programming model and model of parallelism
C-DVM language is an extension of ANSI-C language with annotations named DVM-directives. The directives are implemented as parameters of
DVM(<directive>)
macro, which in sequential program is expanded to empty string.
DVM-directives may be conditionally divided on three subsets:
DVM model of parallelism is based on specific form of data parallelism called SPMD (Single Program, Multiple Data). In this model all the processors concerned execute the same program, but each processor performs its own subset of statements in accordance with the data distribution.
In DVM model at first a user defines multidimensional arrangement of virtual processors, which sections data and computations will be mapped on. The section can be varied from the whole processor arrangement up to a single processor.
Then the arrays to be distributed over processors (distributed data) are determined. These arrays are specified by data mapping directives (section 4). The other variables (distributed by default) are mapped by one copy per each processor (replicated data). A value of replicated variable must be the same on all the processors concerned. Single exception of this rule is variables in parallel loop (section 5.1.3 and 5.1.4).
Data mapping defines a set of local or own variables for each processor. A set of own variables determines the rule of own computations: the processor assigns the values to its own variables only.
DVM model defines two levels of parallelism:
Data parallelism is implemented by distribution of tightly enclosed loop iterations over the processors of the processor arrangement (or the arrangement sections) (see section 5). The loop iteration is executed on one processor entirely. The statements located outside of the parallel loop are executed according to the rule of own computations (see section 5.2).
Task parallelism is implemented by the distribution of data and computations over disjoined sections of processor arrangement (see section 7).
When calculating the value of own variable, the processor may need in values of both own and other (remote) variables. All remote variables must be specified in remote data access directives (see section 6).
The syntax of DVM-directives is described using a Backus-Nour grammar and the following notations:
| ::= | is by definition, |
| | | an alternative construct, |
| [ x ] | optional element/construct, |
| [ x ] | repeat 0 or more times |
| x | repeat 1 or more times; it is equivalent to x [ x ] |
Syntax of the directive.
| directive | ::= DVM ( DVM-directive [ ; DVM-directive ] ) | |
| DVM-directive | ::= specification-directive | executable-directive |
|
| specification-directive | ::= processors-directive | align-directive | distribute-directive | template-directive | shadow-directive | shadow-group-directive | reduction-group-directive | remote-group-directive | task-directive |
|
| executable-directive | ::= realign-directive | redistribute-directive | create-template-directive | parallel-directive | remote-access-directive | create-shadow-group-directive | shadow-start-directive | shadow-wait-directive | reduction-start-directive | reduction-wait-directive | prefetch-directive | reset-directive | map-directive | task-region-directive | on-directive |
|
Constraints:
3 Virtual processor arrangements. PROCESSORS directive
The PROCESSORS directive declares one or more rectangular virtual processor arrangements on the current set of real processors.
Syntax.
| processors-directive | ::= PROCESSORS |
It is possible to define several virtual processor arrangements of different shape if the number of processors in every arrangement will be equal to the number of real processors. If two virtual processor arrangements have the same shape, then corresponding elements of the arrangements are referred to the same processor. Built-in function NUMBER_OF_PROCESSORS () can be used to determine the number of real processors, provided to a program (or subtask).
Note 1. Processor arrangement should be declared as C-array of void * type.
Note 2. To compile program as sequential it should contain the following line:
#define NUMBER_OF_PROCESSORS() 1
Example 3.1. Description of virtual processor arrangements.
DVM(PROCESSORS) void *P[N]; DVM(PROCESSORS) void *Q[NUMBER_OF_PROCESSORS()], *R[2][NUMBER_OF_PROCESSORS()/2];
The value N has to be equal to the value of the function NUMBER_OF_PROCESSORS ( ), and both have to be even.
C-DVM supports distribution by blocks (of equal or different size) and distribution via alignment.
4.1 DISTRIBUTE and REDISTRIBUTE directives
Syntax.
| distribute-directive | ::= | DISTRIBUTE [ dist-directive-stuff ] |
| redistribute-directive | ::= | REDISTRIBUTE distributee dist-directive-stuff [ NEW ] |
| distributee | ::= | array-name |
| dist-directive-stuff | ::= | dist-format... [ dist-onto-clause ] |
| dist-format | ::= | | | |
[BLOCK] [GENBLOCK ( block-array-name )] [WGTBLOCK ( block-array-name,nblock )] [] |
| dist-onto-clause | ::= | ONTO dist-target |
| dist-target | ::= | processors-name [ section-subscript ] |
| section-subscript | ::= | [ subscript [ : subscript ] ] |
Constraints:
Let us consider distribution formats for one dimension of the array (one-dimensional array A[N]) and for one dimension of the processor arrangement (one-dimensional array R[P]). Multi-dimensional distributions are considered in section 4.1.4.
An array dimension is distributed succesively by blocks of [N/P] elements. If N is not divisible by P there will be one partially filled block and may be some empty blocks.
Example 4.1. Distribution by format BLOCK.
| A | B | C | ||||
| R[0] | 0 | 0 | 0 | |||
| 1 | 1 | 1 | ||||
| DVM(PROCESSORS) void * R[4]; | 2 | 2 | ||||
| DVM(DISTRIBUTE [BLOCK] ONTO R) | 3 | |||||
| float A [12], B[13], C[5]; | ||||||
| R[1] | 3 | 4 | 2 | |||
| 4 | 5 | 3 | ||||
| 5 | 6 | |||||
| 7 | ||||||
| R[2] | 6 | 8 | 4 | |||
| 7 | 9 | |||||
| 8 | 10 | |||||
| 11 | ||||||
| R[3] | 9 | 12 | ||||
| 10 | ||||||
| 11 | ||||||
Distribution by blocks of different sizes allows affecting on processor loading balance for algorithms performing different volume of computations for different parts of data area.
Let NB[P] - be an integer array. The following directive
DVM(DISTRIBUTE [ GENBLOCK(NB) ] ONTO R) float A[N];
splits array A on P blocks. The block i of size NB[i] is mapped on processor R[i].
Here:
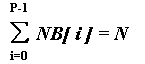
Example 4.2. Distribution by blocks of different size.
| A | ||
| R[0] | 0 | |
| DVM(PROCESSORS) void * R[ 4]; | 1 | |
| int BS[4]={2,4,4,2}; | ||
| DVM(DISTRIBUTE [ GENBLOCK(BS)] ONTO R) | R[1] | 2 |
| float A[12]; | 3 | |
| 4 | ||
| 5 | ||
| R[2] | 6 | |
| 7 | ||
| 8 | ||
| 9 | ||
| R[3] | 10 | |
| 11 | ||
The WGTBLOCK format specifies distribution by blocks according to their relative "weights".
Let WGTBLOCK(WB, NBL) format is specified.
WB(i) defines weight of i-th block for 0£ i £ NBL-1. The blocks are distributed on P processors with balancing of sums of block weights on every processor. The condition
P £ NBL
must be satisfied.
The processor weight is defined as a sum of weights of all the blocks distributed on it . The array dimension is distributed proportionally to processor weights.
BLOCK format is special space of WGTBLOCK(WB,P) format, where WB(i) = 1 for 0£ i £ P-1 and NBL = P.
GENBLOCK format is special case of WGTBLOCK format with some precision.
The example 4.2 can be rewritten using WGTBLOCK format in the following way.
Example 4.3. Distribution by blocks according to weights.
| DVM(PROCESSORS) void * R[ 4]; |
| int WS[12]={2.,2.,1.,1.,1.,1.,1.,1.,1.,1.,2.,2.}; |
| DVM(DISTRIBUTE [ WGTBLOCK(WS,12)] ONTO R) |
| float A[12]; |
In the example 4.3 P = 4 and distribution is identical to the example 4.2.
As distinct from distribution by non-equal blocks, distribution by WGTBLOCK format can be performed for any number of processors from range 1 £ P £ NBL. For given example the size of processor array R can be varied from 1 to 12.
Format [] means, that a dimension is not distributed, that is localized on each processor wholly.
4.1.5 Multidimensional distributions
For multidimensional distributions mapping format is specified for each dimension. The following correspondence is established between dimensions of the array to be distributed and the processor arrangement.
Let the processor arrangement has n dimensions. Number the dimensions not formatted as [ ] from left to right d1, ..., dk. Then dimension di will be mapped on i-th dimension of processor arrangement. The condition k£ n must be satisfied.
Example 4.3. One-dimensional distribution.
| DVM(PROCESSORS) void R1[2]; |
Blocks A |
Processors |
||
| DVM(DISTRIBUTE [BLOCK][] ONTO R1) | 1 | 0 | 1 | |
| float A[100][100]; | 2 | 1 | 2 | |
Example 4.4. Two-dimensional distribution.
| DVM(PROCESSORS) void *R2 [2][2]; | Blocks A | Processors | |||
|
0 |
1 |
||||
| DVM(DISTRIBUTE [BLOCK][BLOCK] | 1 | 2 | 0 | 1 | 2 |
| ONTO R2) float A[100][100]; | 3 | 4 | 1 | 3 | 4 |
4.2 Multidimensional dynamic arrays
4.2.1 Dynamic arrays in C program
DVM system provides extended features to use dynamic arrays. It allows to create multidimensional arrays which sizes along all dimensions are calculated at runtime. (C language allows this for the first dimension only). Multidimensional arrays of different sizes (and sometimes even of different rank) may be passed as parameters. There are two different ways to deal with multidimensional dynamic arrays in C-DVM language. The first one is based on usage of C-style multidimensional arrays; the second one is to simulate multidimensional arrays using one-dimensional ones.
4.2.2 Using C-style multidimensional arrays
The following discipline of declaration, creation, destruction and usage of multidimensional dynamic array elements is provided:
| elem_type | - element type, |
| arrid | - name (identifier) of the array, |
| dimi | - size of i-th dimension of the
array. |
The C-DVM compiler translates these constructions to correct descriptions and Run-Time Library calls. But to translate the program as sequential by ordinary C compiler, the sizes of all the array dimensions dimi (except the first one) must be constants. It is proposed to redefine temporarily them as constants (#define dimi const #undef dimi). The converter removes these redefinitions, more precisely, moves them to the file beginning. Of course, actual ("calculated" in variables) dimensions in case of sequential execution must be equal to selected constant values (see example of Red-black SOR program in Appendix 1).
Example of declaration.
DVM(DISTRIBUTE [BLOCK][BLOCK]) float (*C)[N]; /* two-dimensional distributed array */
4.2.3 Simulation of multidimensional arrays via one-dimensional ones
To operate with multidimensional dynamic arrays in C language other method is used. Memory area (one-dimensional) of required size is created and multidimensional array is simulated by macros arrid(ind1,...,indn) similar to Fortran notation. The macros calculates position of element with specified indexes in the memory area, assigned to the array.
C-DVM converter supports this method also. The array is
Example of array declaration (one-dimensional for sequential execution and two-dimensional for parallel execution) and references to its elements:
int Cdim2; /* The second dimension of array C */ DVM (DISTRIBUTE [BLOCK][]) float *C; #define C(x,y) C[(x)*Cdim2+(y)] . . . C(i,j) = (C(i,j-1)+C(i,j+1))/2;
Aligning array A with distributed array B brings in accordance to each element of A an element or a section of array B. When distributing array B the array A will be distributed simultaneously. If element of B is mapped on the processor, then the element of A, corresponding to element B via alignment, will be also mapped on the same processor.
Method of mapping via alignment performs the following two functions.
4.3.1 ALIGN and REALIGN directives
The following directives describe array aligning:
| align-directive | ::= ALIGN [ align-directive-stuff ] |
| realign-directive | ::= REALIGN alignee align-directive-stuff [ NEW ] |
| align-directive-stuff | ::= align-source... align-with-clause |
| alignee | ::= array-name |
| align-source | ::= [] |
| | [ align-dummy ] | |
| align-dummy | ::= scalar-int-variable |
| align-with-clause | ::= WITH align-spec |
| align-spec | ::= align-target [ align-subscript ] |
| align-target | ::= array-name |
| | template-name | |
| align-subscript | ::= [ int-expr ] |
| | [ align-subscript-use ] | |
| | [] | |
| align-subscript-use | ::= [ primary-expr * ] align-dummy [ add-op primary-expr ] |
| primary-expr | ::= int-constant |
| | int-variable | |
| | ( int-expr ) | |
| add-op | ::= + |
| | - |
Constraints:
Let the alignment of two arrays is specified by the directive
DVM(ALIGN [d1] [dn] WITH B[ard1] [ardm]) float A[100] [100];
where:
di - specification of i-th
dimension of aligned array A,
ardj - specification of j-th
dimension of base array B.
If di is specified by integer variable I, one and only one dimension of array B, specified by linear function ardj = a*I + b must exist. Therefore, the number of array A dimensions, specified by identifiers (align-dummy) must be equal to the number of array B dimensions, specified by the linear function.
Let i-th dimension of array A has the size Ai, and j-th dimension of array B, specified by linear function a*I + b, has the size Bj. Since the parameter I is defined on the value set 0 Ai-1, the following conditions must be satisfied:
b ³ 0; a*(Ai 1)+ b < Bj
If di is empty, the i-th dimension of array A will be local on each processor independently from array B distribution (it is analogue of local dimension in DISTRIBUTE directive).
If ardi is empty, the array A will be replicated along the j-th dimension of the array B (it is analogue of partial replication on the processor arrangement).
If ardi = int-expr, the array A is aligned with the section of the array B.
Example 4.7. Aligning arrays
DVM(DISTRIBUTE [BLOCK][BLOCK]) float B[10][10]; DVM(DISTRIBUTE [BLOCK]) float D[20]; /* aligning the vector with the first line of B) */ DVM(ALIGN [i] WITH B[1][i] ) float A[10]; /* replication of the vector aligning it with each line */ DVM(ALIGN [i] WITH B[][i]) float F[10]; /* collaps: each matrix column corresponds to the vector element */ DVM(ALIGN [][i] WITH D[i]) float C[20][20]; /* alignment using stretching */ DVM(ALIGN [i] WITH D[2*i]) float E[10]; /* alignment using rotation and stretching */ DVM(ALIGN [i][j] WITH C[2*j][2*i] ) float H[10,10];
Let the sequence of alignments A f1 B f2 C, be specified; f2 be the alignment of the array B with the array C , and f1 be the alignment of the array A with the array B. The arrays A and B are considered as aligned with array C by definition. The array B is aligned by function f2 directly and array A is aligned by composite function f1(f2) indirectly. Therefore applying REALIGN directive to the array B does not cause redistribution of array A.
Generally a set of ALIGN specifications is a set of trees. At that every tree root must be distributed by DISTRIBUTE or REDISTRIBUTE directives. When REDISTRIBUTE directive is executed, the whole alignment tree is redistributed.
4.3.2 TEMPLATE and CREATE_TEMPLATE directives
If values of linear function a*I + b are beyond base array dimension, it is necessary to define a dummy array - referred to as an alignment template using the following directive.
| template-directive | ::= TEMPLATE [ size ] |
Then it is necessary to align both arrays with the template. The template is defined using DISTRIBUTE and REDISTRIBUTE directives. The template elements don't require real memory. They specify the processor the elements of aligned arrays must be mapped on.
Consider the following example.
Example 4.8. Aligning with template.
DVM(DISTRIBUTE [BLOCK]; TEMPLATE[102]) void * TABC; DVM(ALIGN B[i] WITH TABC[i]) float B[100]; DVM(ALIGN A[i] WITH TABC[i+1]) float A[100]; DVM(ALIGN C[i] WITH TABC[i+2]) float C[100]; . . . DO(i ,1, 98,1) A[i] = C[i-1] + B[i+1];
If the sizes of template (and of arrays) are unknown statically the following executable directive should be used:
| create-template-directive | ::= CREATE_TEMPLATE template_name size |
Example 4.9. Aligning dynamic arrays with template.
DVM(DISTRIBUTE [BLOCK]; TEMPLATE ) void *TABC; DVM(ALIGN B[i] WITH TABC[i]) float *B; DVM(ALIGN A[i] WITH TABC[i+1]) float *A; DVM(ALIGN C[i] WITH TABC[i+2]) float *C; int N; . . . N= ... /* size calculation */...; DVM(CREATE_TEMPLATE TABC[N]); A=malloc(N*sizeof(float)); B=malloc(N*sizeof(float)); C=malloc(N*sizeof(float)); . . . DO(i ,1, N-2,1) A[i] = C[i-1] + B[i+1];
If DISTRIBUTE or ALIGN directive is not specified for the data, then the data are allocated on each processor (full replication). The same distribution can be specified by DISTRIBUTE directive with [] format for each dimension. But in this case access to the data will be less effective.
| C-DVM - contents | Part 1(1-4) | Part 2 (5-11) | Part 3 (Appendixes) |